Biokemi keywords samlet PDF

| Title | Biokemi keywords samlet |
|---|---|
| Author | Siw Henrichsen |
| Course | Biokemi |
| Institution | Danmarks Tekniske Universitet |
| Pages | 29 |
| File Size | 776.4 KB |
| File Type | |
| Total Downloads | 2 |
| Total Views | 73 |
Summary
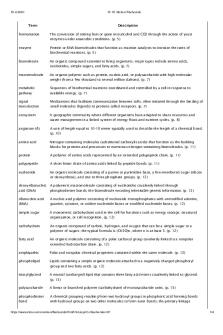
Term Descriptionfermentation The conversion of rotting fruit or grain into alcohol and CO2 through the action of yeast enzymes under anaerobic conditions. (p. 5)enzyme Protein or RNA biomolecules that function as reaction catalysts to increase the rates of biochemical reactions. (p. 5)biomolecule An...
Description
16.4.2020
W. W. Norton Flashcards
Term
Description
fermentation
The conversion of rotting fruit or grain into alcohol and CO2 through the action of yeast enzymes under anaerobic conditions. (p. 5)
enzyme
Protein or RNA biomolecules that function as reaction catalysts to increase the rates of biochemical reactions. (p. 5)
biomolecule
An organic compound essential to living organisms; major types include amino acids, nucleotides, simple sugars, and fatty acids. (p. 7)
macromolecule
An organic polymer such as protein, nucleic acid, or polysaccharide with high molecular weight (from a few thousand to several million daltons). (p. 7)
metabolic pathway
Sequences of biochemical reactions coordinated and controlled by a cell in response to available energy. (p. 7)
signal transduction
Mechanisms that facilitate communication between cells, often initiated through the binding of small molecules (ligands) to proteins called receptors. (p. 7)
ecosystem
A geographic community where different organisms have adapted to share resources and waste management in a linked system of energy flows and nutrient cycles. (p. 8)
angstrom (Å)
A unit of length equal to 10-10 meter typically used to describe the length of a chemical bond. (p. 10)
amino acid
Nitrogen-containing molecules (substituted carboxylic acids) that function as the building blocks for proteins and precursors to numerous nitrogen-containing biomolecules. (p. 11)
protein
A polymer of amino acids represented by an extended polypeptide chain. (p. 11)
polypeptide
A short linear chain of amino acids linked by peptide bonds. (p. 11)
nucleotide
An organic molecule consisting of a purine or pyrimidine base, a five-membered sugar (ribose or deoxyribose), and one to three phosphate groups. (p. 12)
deoxyribonucleic acid (DNA)
A polymeric macromolecule consisting of nucleotides covalently linked through phosphodiester bonds; the biomolecule encoding inheritable genetic information. (p. 12)
ribonucleic acid (RNA)
A nucleic acid polymer consisting of nucleoside monophosphates with unmodified adenine, guanine, cytosine, or uridine nucleotide bases or modified nucleotide bases. (p. 12)
simple sugar
A monomeric carbohydrate used in the cell for functions such as energy storage, structural organization, or cell recognition. (p. 12)
carbohydrate
An organic compound of carbon, hydrogen, and oxygen that can be a simple sugar or a polymer of sugars; the typical formula is (CH2O)n, where n is at least 3. (p. 12)
fatty acid
An organic molecule consisting of a polar carboxyl group covalently linked to a nonpolar extended hydrocarbon chain. (p. 12)
amphipathic
Polar and nonpolar chemical properties contained within the same molecule. (p. 12)
phospholipid
Lipids containing a simple organic molecule attached to a negatively charged phosphoryl group and two fatty acids. (p. 12)
triacylglycerol
A neutral (uncharged) lipid that contains three fatty acid esters covalently linked to glycerol. (p. 13)
polysaccharide
A linear or branched polymer (carbohydrate) of monosaccharide units. (p. 13)
phosphodiester bond
A chemical grouping resulting from two hydroxyl groups in phosphoric acid forming bonds with hydroxyl groups on two other molecules to form ester bonds; the primary linkage
https://wwnorton.com/common/flashcards/FrmPrint.aspx?s=biochem&c=01
1/4
16.4.2020
W. W. Norton Flashcards
between nucleotides in DNA and RNA. (p. 13) peptide bond
A covalent bond between the α amino group of one amino acid and the α carboxyl group of another amino acid. (p. 14)
amylose
An α-1,4 linear form of starch. (p. 14)
chitin
A linear polysaccharide consisting of N-acetylglucosamine units linked by β-1,4 glycosidic bonds. Chitin is a major component of insect and crustacean exoskeletons and fungal cell walls. (p. 14)
metabolite
Any of a group of small biomolecules that serve as reactants and products in biochemical reactions within cells. (p. 15)
systems biology
The study of complex chemical reaction networks in cells. (p. 15)
metabolic flux
The rate at which reactants and products are interconverted in a metabolic pathways. (p. 15)
linear metabolic
A metabolic pathway in which each reaction generates only a single product, which is a
pathway
reactant for the next reaction in the pathway. (p. 17)
forked pathway
A metabolic pathway that generates two products, each of which undergoes a different metabolic fate. (p. 17)
cyclic pathway
A metabolic pathway containing several metabolites that regenerate during each turn of the cycle, serving as both reactants and products. (p. 17)
prokaryote
A single-celled organism that lacks a nucleus and other membrane-bound organelles; this category includes all bacteria. (p. 17)
eukaryote
A cell that contains a nucleus and other organelles bounded by membranes, creating microenvironments for biochemical reactions. (p. 17)
capsule
An outer layer, usually of polysaccharides, that surrounds some prokaryotic cells, particularly bacteria. (p. 17)
cytoplasm
A cell's contents within the plasma membrane (but not including the nucleus in eukaryotic cells), including organelles; the site of most cellular activities. (p. 17)
chromosome
A DNA molecule that functions to store and transmit genetic information. (p. 17)
nucleoid
In bacteria, a region containing the chromosome (without any surrounding membrane). (p. 17)
flagellum
An extracellular structure used for cell movement by bacteria and sperm cells. (p. 18)
pilus
A hair-like appendage on the surface of bacteria, used in cell movement and reproduction. (p. 18)
plasmid
A circular DNA molecule that replicates independently of the chromosome; bacterial plasmids encode genes for cell mating, antibiotic resistance, and pathogenesis. (p. 18)
genome
All of the genetic information in a cell or virus contained in DNA or RNA. (p. 18)
chromatin
In eukaryotes, a complex of DNA and proteins that constitutes chromosomes. (p. 18)
nucleus
In eukaryotic cells, a membrane-bound organelle that contains chromosomes. (p. 18)
nucleolus
In eukaryotic cells, a part of the nucleus where ribosomes are assembled from ribosomal RNA and protein. (p. 18)
ribosome
Large RNA-protein complexes that mediate protein synthesis in prokaryotic and eukaryotic cells. (p. 18)
cytoskeleton
In eukaryotic cells, a network of intracellular filaments, consisting of oligomeric proteins, that maintains cell structure. (p. 19)
https://wwnorton.com/common/flashcards/FrmPrint.aspx?s=biochem&c=01
2/4
16.4.2020
W. W. Norton Flashcards
mitochondrion pl. mitochondria
The eukaryotic organelle responsible for many of the metabolic reactions involved in energy conversion and production of ATP. (p. 19)
peroxisome
In eukaryotic cells, an organelle containing enzymes for forming or destroying peroxides. (p. 19)
lysosome
In eukaryotic cells, a membrane-bound organelle involved in degradation and detoxification of macromolecules. (p. 19)
chloroplast
In plant cells, an organelle that converts light energy into chemical energy. (p. 19)
vacuole
A membrane-bound organelle in many types of eukaryotic cells, particularly plant cells, that stores metabolites and also isolates molecules that might be harmful to the cell. (p. 19)
endoplasmic
In eukaryotic cells, highly invaginated membrane structures that sequester ribosomes for
reticulum
protein synthesis. (p. 19)
Golgi apparatus
In eukaryotic cells, a membranous structure required for protein translocation within the cell and in facilitating protein secretion at the plasma membrane. (p. 19)
microtubule
Cable-like component of the cytoskeleton that enables an animal cell to move by extending the plasma membrane in one direction while retracting it at the opposite end of the cell. (p. 19)
endosymbiotic
The theory proposing that eukaryotic cells evolved about 1.5 billion years ago as a result of
theory
large predatory cells engulfing aerobic bacteria or cyanobacteria, giving rise to mitochondria and chloroplasts, respectively. (p. 19)
ligand
A small molecule that is often a metabolite, hormone, or peptide and which binds to target proteins and alters their structure and function to control biochemical processes. (p. 20)
base pair
Two nucleotides in nucleic acid chains that are paired by hydrogen bonding, such as C-G, A-T, or A-U. (p. 24)
base stacking
Stabilizing interactions between the aromatic rings of the nucleotide bases within the interior of the DNA helix. (p. 25)
central dogma
The description of information transfer in molecular biology, which flows from DNA to RNA to proteins. (p. 25)
DNA replication
The enzyme-mediated process of doubling the DNA content of a cell during division. (p. 26)
gene
A segment of DNA that codes for a transcribed RNA molecule; the functional unit of heredity. (p. 26)
DNA transcription
The process of generating RNA from a DNA template. (p. 26)
messenger RNA (mRNA)
A molecule of RNA that serves as a template for protein synthesis. (p. 26)
mRNA translation
The process by which a ribosome decodes a molecule of mRNA and synthesizes a corresponding protein. (p. 26)
small nuclear RNA (snRNA)
Small RNA molecules involved in RNA processing. (p. 26)
micro RNA (miRNA)
Genome encoded small RNA molecules that regulate gene expression and mRNA translation. (p. 26)
ribosomal RNA (rRNA)
A type of RNA that forms the major component of ribosomes. (p. 26)
transfer RNA (tRNA)
The adaptor molecule in protein synthesis that delivers an amino acid to the ribosome. (p. 26)
https://wwnorton.com/common/flashcards/FrmPrint.aspx?s=biochem&c=01
3/4
16.4.2020
W. W. Norton Flashcards
reverse transcription
The process that converts RNA into DNA, most often related to replication of retroviruses. (p. 27)
transcriptome
The collection of DNA transcripts (RNA products) generated by DNA transcription. (p. 27)
proteome
The complete collection of proteins in a cell or organism. (p. 27)
template strand
A DNA sequence that has the opposite 5′ to 3′ polarity as the corresponding mRNA transcript; it is complementary to the DNA coding strand. (p. 27)
coding strand
A DNA sequence that has the same 5′ to 3′ polarity as the corresponding mRNA transcript; it is complementary to the DNA template strand. (p. 27)
natural selection
The change in the frequency of genes in a population under conditions that favor some genes over others. (p. 28)
wild-type
A fully functional protein-coding sequence without any mutations. (p. 28)
germ-line cell
Cells that give rise to an egg or sperm cell in eukaryotes. (p. 28)
somatic cell
Any cell in an organism other than a germ-line cell. (p. 28)
bioinformatics
The use of computational tools to probe and analyze large data sets of biological information, typically whole genomes or proteomes. (p. 29)
orthologous
One of a set of highly conserved gene sequences that arose from a common ancestral gene
gene
and encode proteins with the same function in different species. (p. 31)
gene duplication
A mechanism by which duplication of a region of DNA containing a gene can lead to evolution of new genetic material. (p. 32)
paralogous
Highly conserved genes within the same species; most often derived from the process of gene
genes
duplication. (p. 32)
https://wwnorton.com/common/flashcards/FrmPrint.aspx?s=biochem&c=01
4/4
16.4.2020
W. W. Norton Flashcards
Term bioenergetics
Description Energy conversion processes in biological systems, including transformation of solar energy into chemical energy and interconversion of chemical energy through oxidation and reduction of organic molecules. (p. 40)
homeostasis
The use of energy to maintain a dynamic steady state of an organism that can adjust to changing environmental conditions. (p. 40)
equilibrium
The state of a system in which no net change occurs. (p. 40)
solar energy
Energy produced through thermonuclear reactions in the Sun. (p. 41)
photosynthetic autotroph
An organism that can use photosynthesis to oxidize water and produce oxygen, generating chemical energy in the form of glucose. (p. 41)
aerobic respiration
A set of metabolic processes and reactions that uses oxygen to generate ATP. (p. 41)
heterotroph
An organism that cannot convert solar energy to chemical energy directly, but must depend on nutrients obtained from autotrophs and other heterotrophs as a source of energy. (p. 41)
photosynthesis
The use of solar energy to oxidize water, capture chemical energy, and generate oxygen. (p. 41)
carbon fixation
The conversion of carbon dioxide into other organic compounds, particularly glucose; often considered as part of photosynthesis. (p. 42)
biosphere
All the living organisms on Earth, considered as a whole. (p. 42)
redox reaction
An oxidation–reduction reaction in which electrons are transferred from a compound of lower reduction potential (more negative) to one of higher reduction potential (more positive) as in the electron transport system. (p. 42)
photooxidation
Oxidation caused by light; used particularly for oxidation of chlorophyll, resulting in the transfer of an electron from chlorophyll to an acceptor molecule such as pheophytin. (p. 42)
photophosphorylation
The conversion of ADP to ATP coupled to the transfer of electrons in photosynthesis. (p. 42)
oxidative phosphorylation
A metabolic pathway that oxidizes nutrients, particularly glucose, to generate ATP from ADP. (p. 42)
first law of thermodynamics
In any physical or chemical change, the total amount of energy in the universe remains the same, even though the form of energy may change. (p. 44)
second law of thermodynamics
In the absence of an energy input, all spontaneous processes in the universe tend toward dispersal of energy (disorder), and moreover, the measure of this disorder, called entropy, is always increasing in the universe. (p. 44)
entropy
A measure of the spreading of energy; also a measure of the disorder (randomness) in a system. (p. 44)
enthalpy (H)
The heat content of a system. (p. 44)
bomb calorimeter
A device in which a compound is combusted by a spark at constant volume in the presence of pure oxygen (completely oxidized); the amount of heat exchanged between the reaction chamber and a surrounding water jacket is a measure of enthalpy change. (p. 44)
exothermic
A reaction that releases heat and ΔH < 0. (p. 45)
https://wwnorton.com/common/flashcards/FrmPrint.aspx?s=biochem&c=02
1/4
16.4.2020
W. W. Norton Flashcards
endothermic
A reaction that absorbs heat and ΔH > 0. (p. 45)
calorie (cal)
A unit of energy; the amount of heat energy required to raise 1 gram of water by 1 °C using a calorimeter; a Calorie is 103 calories or 1 kilocalorie (kcal), which is equal to 4.184 kilojoules (kJ). (p. 45)
joule (J)
The SI unit of energy; the amount of work done (or energy transferred) when a force of 1 Newton displaces an object by 1 meter in the direction of the force. (p. 45)
Gibbs free energy (G)
A measure of the spontaneity of a reaction, defined as the difference between the enthalpy and entropy of a system at a given temperature (G = H - TS). (p. 48)
exergonic
A chemical reaction that releases energy and is favorable in the direction written with a ΔG < 0. (p. 48)
endergonic
A chemical reaction that requires energy and is unfavorable in the direction written with a ΔG > 0. (p. 48)
standard free energy change (ΔG°)
A reference point for comparing chemical reactions under a defined set of conditions (1 atm pressure, 298 K, 1 M concentration of reactants and products): ΔG° = ΔH° - TΔS° (p. 48)
biochemical standard
Reaction conditions of 1 atm pressure, 298 K temperature, 1 M concentrations of
conditions
reactants and products, pH of 7, and concentration of water at 55.5 M. (p. 48)
equilibrium constant
A measure of the directionality of a reaction under standard conditions, where all
(Keq)
products and reactants start at 1 M and proceed to their equilibrium concentrations. (p. 49)
adenylate system
A group of several phosphoryl transfer reactions that interconvert ATP, ADP, and AMP. (p. 53)
isozymes
Functionally related enzymes encoded by different genes. (p. 54)
isoforms
Functionally distinct proteins transcribed from the same gene. (p. 54)
energy charge (EC)
A measure of the energy state of a cell in terms of ATP, ADP, and AMP ratios: EC = [ATP] + 0.5[ADP] / [ATP] + [ADP] + [AMP] (p. 54)
catabolic pathway
A metabolic pathway that converts energy-rich compounds into energy-depleted compounds, releasing energy for the cell. (p. 55)
anabolic pathway
A metabolic pathway for the biosynthesis of biomolecules from smaller precursors. (p. 55)
hydrogen bond
A weak noncovalent bond in which hydrogen is shared between two electronegative atoms. (p. 57)
proton hopping
A series of hydrogen-bond exchanges between adjacent H2O molecules leading to the transient formation of hydronium ions (H3O+). (p. 58)
ionic interaction
A weak interaction between oppositely charged atoms or groups. (p. 61)
van der Waals
A weak interaction between the dipoles of nearby electrically neutral molecules. (p...
Similar Free PDFs
Biokemi keywords samlet
- 29 Pages
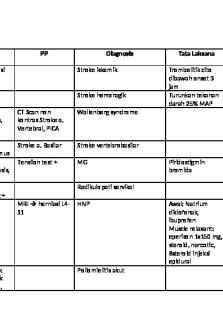
Keywords UKMPPD
- 43 Pages

Samlet noter
- 44 Pages

Romanticism- Keywords
- 3 Pages

BUS201 Keywords
- 5 Pages

Materialelære samlet besvarelser
- 99 Pages

Biokemi - Begreber + Opsummeringer
- 10 Pages
![[07.1] Live de keywords](https://pdfedu.com/img/crop/172x258/zwrgkope9k31.jpg)
[07.1] Live de keywords
- 82 Pages

Keywords by Raymond Williams
- 2 Pages

Topics and Keywords
- 2 Pages

Materialelære samlet noter og opgaver
- 109 Pages

Keywords of art history terms
- 2 Pages

TFG Citacion y uso de keywords
- 10 Pages
Popular Institutions
- Tinajero National High School - Annex
- Politeknik Caltex Riau
- Yokohama City University
- SGT University
- University of Al-Qadisiyah
- Divine Word College of Vigan
- Techniek College Rotterdam
- Universidade de Santiago
- Universiti Teknologi MARA Cawangan Johor Kampus Pasir Gudang
- Poltekkes Kemenkes Yogyakarta
- Baguio City National High School
- Colegio san marcos
- preparatoria uno
- Centro de Bachillerato Tecnológico Industrial y de Servicios No. 107
- Dalian Maritime University
- Quang Trung Secondary School
- Colegio Tecnológico en Informática
- Corporación Regional de Educación Superior
- Grupo CEDVA
- Dar Al Uloom University
- Centro de Estudios Preuniversitarios de la Universidad Nacional de Ingeniería
- 上智大学
- Aakash International School, Nuna Majara
- San Felipe Neri Catholic School
- Kang Chiao International School - New Taipei City
- Misamis Occidental National High School
- Institución Educativa Escuela Normal Juan Ladrilleros
- Kolehiyo ng Pantukan
- Batanes State College
- Instituto Continental
- Sekolah Menengah Kejuruan Kesehatan Kaltara (Tarakan)
- Colegio de La Inmaculada Concepcion - Cebu
